
Debra Knisley's Home Webpage

Professor of Mathematics
Director of the Institute for
Quantitative Biology
Gilbreath Hall Room 307B
knisleyd@etsu.edu
423) 439-6975 |
|
Director of the Institute for Quantitative Biology:
Building upon a rich history of collaboration between the Department of
Mathematics and Statistics and the Department of Biological Sciences, the
Institute promotes interdisciplinary research and instruction at the interface
of computational and life sciences. Our membership spans six colleges: Arts and
Sciences, Business and Technology, Medicine, Nursing, Pharmacy and Public
Health.
see www.etsu.edu/iqb
Professor of Mathematics:
The Department of Mathematics and Statistics offers a
rich environment for student learning. Students frequently conduct joint
research projects with the faculty, travel to professional meetings where
they present their findings and participate in other activities outside the
classroom. The Department offers both a B.S. and M.S. in Mathematics which
prepares them for a wide range of career choices.
see www.etsu.edu/cas/math
My Research
My research is in the field of Computational Biology. In my research, I use
networks (graph-theoretic models) to model biomolecular structures such as RNA, DNA
and proteins. Using these models, we attempt to quantify the structural,
biophysical and biochemical properties so that we may apply machine learning
techniques to reveal their features. This helps us to identify the roles
these molecules play within the biological systems in which they occur.
Membrane proteins are a class of proteins that are becoming of increasing
importance. We are studying the structure, function and dynamics of the
cystic fibrosis conductance regulator protein, an ATP Binding Cassette
Membrane Protein.
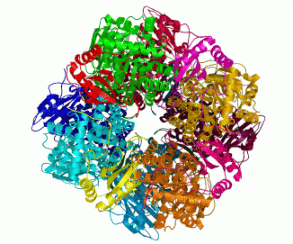
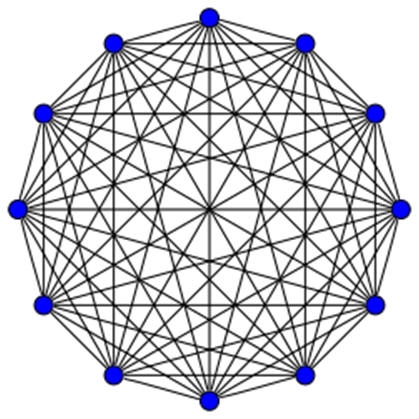
Professional Service
Elected to the Board of Directors for the MidSouth Computational Biology and
Bioinformatics Society- An affiliate of the International Society for
Computational Biology (2012-2015) MCBIOS
Co-Chair of the Organizing Committee for the 25th Annual
Cumberland Conference on Combinatorics, Graph Theory and Computing to be
held May 10-12, 2012
Cumberland25
Students
- Cade Herron: A comparative study of the Human ATP-Binding
Cassette Proteins using graphs
- Chelsea Ross: Molecular descriptors of multigraph representations of
secondary RNA structure
- Cade Herron: Graph-theoretic models of mutations of the nucleotide
binding domain 1 of the cystic fibrosis transmembrane conductance regulator
- Josh Brooks: Network Properties of hierarchical (2,r)-regular graphs
- Russell Sharp: Connectivity properties of alpha-overlap graphs
Former Students
- Alissa Rockney: Classifying Secondary RNA structures using the dual
graph and machine learning
- Sean Milam: Graph-theorectic modeling of the
Oligonucleotide/Oligosaccharide Binding Fold in DNA repair proteins
- Denise Koessler: A predictive model for secondary RNA structure
using graphs and a neural network
SUMMA: Strengthening Underrepresented Minorities in Mathematical Achievement:
ETSU
SUMMA 2008: Discrete Mathematical Models in Molecular Biology:
Solutions for the future
ETSU SUMMA 2007:
Biomolecular Graphs
ETSU SUMMA
2005: Graphs
with applications in biology and computer science
ETSU SUMMA 2004:
Graph-theoretic analysis of RNA
________________________________________________________________________
Education:
B.S. (1976) Tennessee Technological University
M.S. (1979) Tennessee Technological University
Ph.D. (1989) University of Memphis
Recent (selected) Publications:
-
Seeing the results of a mutation with a vertex weighted graph, Knisley D, Knisley J, (to appear in BMC Bioinformatics Supplemental issue- Highlights from the 3rd IEEE Symposium on Biological Data Visualization).
-
Vertex-weighted graphs and their applications, Knisley D. and Knisley J., invited paper (to appear in Utilitas Mathematica)
-
Network properties of nested (t,r)-regular graphs, Knisley D. Brooks J and Knisley J, Journal of Combinatorial Mathematics and Computing, vol 88 (February 2014)
-
Graph-theoretic models of mutations of the nucleotide binding domain 1 of the cystic fibrosis transmembrane conductance regulator, Knisley D, Knisley J and Herron C, Computational Biology Journal, Article # 138969 (2013)
-
Classifying multigraph models of secondary RNA structure using
graph-theoretic descriptors, Knisley D, Knisley J, Rockney A and Ross C,
ISRN Bioinformatics vol 12 (2012), Article ID 157135,
www.isrn.com/journals/bioinformatic/aip/157135
-
Predicting protein-protein interactions using
graphical invariants and a neural network, Knisley J and Knisley D,
Journal
of Computational Biology and Chemistry 35, pp 108-113, (2011)
-
Predicting Flavonoid UGT Regioselectivity,
Jackson R, Knisley D, McIntosh C and Pfeiffer P,
Advances in Bioinformatics,
Vol
2011, Article ID 506583, (2011)
-
On properties of the α-overlap graphs, Godbole A,
Knisley D and Norwood R, Congr. Num. Vol. CCIV, pp 161-172, (2010)
- On the chromatic number of alphabet-overlap graphs, Knisley D, Nigussie
Y and Por A, Journal of Combinatorial Mathematics and Computational
Computing, JCMCC, 73, 3-13, 2010.
-
A predictive model for secondary RNA structure using graph theory and a
neural network, Koessler, D, Knisley, D, Knisley J, Haynes T, BMC
Bioinformatics 2010, 11(Suppl 6):S21 (7 Oct 2010)
- A graph-theoretic model based on primary and predicted secondary
structure reveals functional specificity in a set of
UDP-glucosyltransferases, Knisley D, Seier E, Lamb D, Owens D and McIntosh
C, Proceedings of the 2009 International Conference on Bioinformatics,
Computational Biology, Genomics and Cheminformatics,bcbgc-09, Loging,
Doble, Sun, Malone (Editors) ISBN: 978-1-60651-009-4, 2009
- Using a neural network to identify secondary RNA structures
quantified by graphical invariants, Haynes, T, Knisley D, Knisley J, Communications in Mathematical and Computer Chemistry,
MATCH: 60(2),
277-290, 2008.
- Network Properties of (t,r)-regular graphs, Knisley D,
Knisley J and Williams D., Proceedings of the 2008 International
Conference on Theoretical and Mathematical Foundations of Computer Science,
Zoran, Sipser, Radha, Wei (Editors) ISBN: 978-1-60651-006-3, 2008
- Graph-theoretic models in Chemistry and Molecular Biology, Book
Chapter in the Handbook of Applied Algorithms: Solving Scientific,
Engineering and Practical Problems, Stojmenovic and Nayak
(Editors), John Wiley, 2007.
- A quantitative analysis of secondary RNA structure using
domination based parameters on trees, Haynes T, Knisley D, Seier E and Zou
Y, BMC Bioinformatics 7: (108), 2006.
Institutes: Mathematics-
Institutes: Computational Biology-
Return to Mathematics' home page




